Projects
Expanded Clinical Application of RNA Sequencing
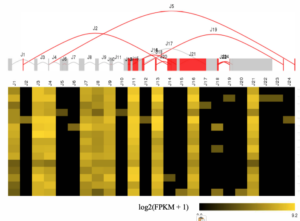
Approximately 95% of clinical molecular oncology tests assess DNA, with only 5% being RNA based. However, RNA-based testing has the potential to both complement and dramatically expand upon DNA-based testing. RNA testing has a higher yield in identification of clinically relevant gene fusions, and it allows examination of splice alterations, gene expression changes, and gene expression profiling signatures. Similar to the shift toward broader DNA-based molecular oncology assays, we expect that over time, we will see a shift from RNA-based assays focusing on a single gene or indication to more comprehensive transcriptome sequencing. Our group, in collaboration with many others at UNC, is drawing upon the extensive institutional experience with RNA sequencing and analysis, from projects such as The Cancer Genome Atlas, to the long-term goal of applying comprehensive transcriptome sequencing in the clinical setting.
Gene Expression Features and Signatures Predicting Tumor Behavior and Response to Therapies
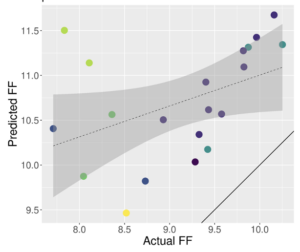
One of our initial projects related to the expanded clinical application of RNA sequencing involves development and application of gene expression features and signatures that predict tumor biology and response to therapies, initially immunotherapies. While immunotherapy regimens generally have a favorable side-effect profiles, they can result in significant treatment-related adverse events, more commonly when given in combinations. Current biomarkers used to guide selection of immunotherapy in clinical practice have limitations, and recently developed RNA sequencing-based gene signatures are promising biomarkers to guide use of immunotherapy in patients with advanced cancer. In collaboration with Drs. Kim, Perou, and Vincent, we are applying existing and novel immune signatures that have been modeled for clinical FFPE tissues. Development efforts are focused broadly on solid tumors with an emphasis on breast and endometrial cancers. Predictive signatures will be evaluated in a retrospective and then prospective clinical trial setting to evaluate clinical validity and utility.
Application of Liquid Biopsy Assays in Cancer
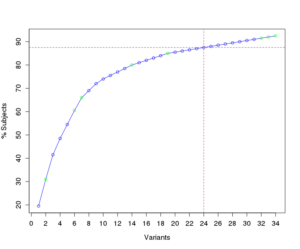
Liquid biopsy refers to a broad category of minimally invasive blood tests that detect cancer cells from a tumor or fragments of tumor-derived nucleic acids or proteins that are in the blood (PMID: 29504847). Our group is focused on the development and application of clinical tests designed to detect somatic variants in cell-free, circulating tumor DNA (ctDNA) and RNA (ctRNA) in multiple areas. Current projects include use of ctDNA and ctRNA for cancer genotyping, advanced cancer monitoring, and early-stage cancer molecular residual disease (MRD) monitoring. The laboratory uses a combination of targeted, exome, and genome sequencing approaches to support this work.
Correlative Genomic Biomarker Studies
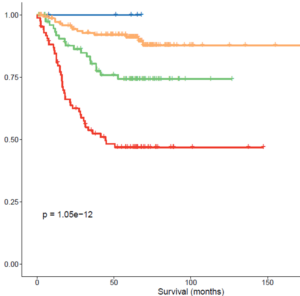
Correlative genomic biomarker studies embedded in clinical trials provide a unique opportunity to rigorously evaluate the clinical validity and clinical utility of genomic assays, which is ultimately needed for a biomarker or test to be used in the clinical setting. With Drs. Hayes and Perou, I direct the Network Group Integrated Translational Science Center at UNC for the NCI National Clinical Trials Network. This Center supports the application of genomics testing, including genome, exome, RNA, ctDNA and single-cell sequencing, to NCI clinical trials. Likewise, Drs. Merker and Perou are Faculty Directors for the Lineberger Comprehensive Cancer Center Translational Genomics Laboratory, a sequencing facility that provides comprehensive cancer genomic services, including to cooperative group and investigator-initiated cancer clinical trials.
Molecular Profiling in Endometrial Cancer
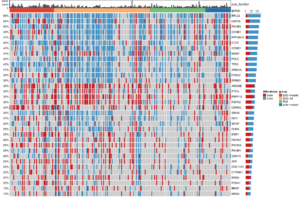
Endometrial cancer, the fourth most common cancer in US women, shows one of the largest racial disparities in US cancers. Furthermore, the frequency and mortality of endometrial cancer continues to rise. Compared to all other ethnic and racial groups, Black women have a >90% higher 5-year mortality rate from endometrial cancer. As part of the multi-disciplinary Carolina Endometrial Cancer Study (CECS) and (Program on Translational Research in Endometrial Cancer Disparities (UNC TREND), our group is focused on better understanding the potential molecular and genetic reasons underlying this disparity. Integrated genomic characterization of endometrial cancer has significantly contributed to our understanding of the molecular and genetic basis of this cancer, enabling the application of molecular classifications with prognostic and treatment implications. A central focus of CECS and UNC TREND is to integrate these molecular analyses into a North Carolina population-based study with clinical and epidemiologic annotations to facilitate endometrial cancer disparities research.
Other Areas of Interest
- Heritable Cancer Predisposition
- Bioinformatics Software Development